Using Databricks Notebooks to run an ETL process

I recently wrote a blog on using ADF Mapping Data Flow for data manipulation. As part of the same project, we also ported some of an existing ETL Jupyter notebook, written using the Python Pandas library, into a Databricks Notebook.
This notebook could then be run as an activity in a ADF pipeline, and combined with Mapping Data Flows to build up a complex ETL process which can be run via ADF. Databricks is built on Spark, which is a "unified analytics engine for big data and machine learning". It allows you to run data analysis workloads, and can be accessed via many APIs (Scala, Java, Python, R, SQL, and now .NET!), though only Scala, Python and R are currently built into Notebooks. This means that you can build up data processes and models using a language you feel comfortable with.
Connecting the Databricks account to ADF
To start with, you create a new connection in ADF. By choosing compute, and then Databricks, you are taken through to this screen:
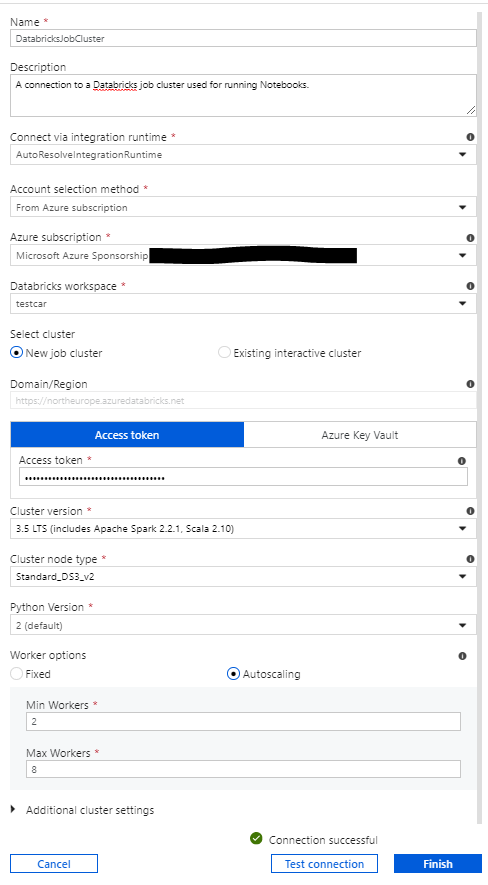
Here you choose whether you want to use a job cluster or an existing interactive cluster. If you choose job cluster, a new cluster will be spun up for each time you use the connection (i.e. each time you run a notebook). It should be noted that cluster spin up times are not insignificant - we measured them at around 4 minutes. Therefore, if performance is a concern it may be better to use an interactive cluster. An interactive cluster is a pre-existing cluster. These can be configured to shut down after a certain time of inactivity. This is also an excellent option if you are running multiple notebooks within the same pipeline. Using job clusters, one would be spun up for each notebook. However, if you use an interactive cluster with a very short auto-shut-down time, then the same one can be reused for each notebook and then shut down when the pipeline ends. However, you pay for the amount of time that a cluster is running, so leaving an interactive cluster running between jobs will incur a cost.
Creating the Notebook
Once the Databricks connection is set up, you will be able to access any Notebooks in the workspace of that account and run these as a pipeline activity on your specified cluster. You can either upload existing Jupyter notebooks and run them via Databricks, or start from scratch. As I've mentioned, the existing ETL notebook we were using was using the Pandas library. This is installed by default on Databricks clusters, and can be run in all Databricks notebooks as you would in Jupyter. However, the data we were using resided in Azure Data Lake Gen2, so we needed to connect the cluster to ADLS.
This was done using a secret which can be created using the CLI as follows:
- Create a personal access token in the "Users" section in Databricks
- Install the Databricks CLI using pip with the command
pip install databricks-cli - Create a "secret" in the Databricks account
To create the secret use the command databricks configure --token, and enter your personal access token when prompted. Create a scope for that secret using:
Databricks secrets create-scope --scope <name of scope> --initial-manage-principal users
initial-manage-principal must be set to "users" when using a non-premium-tier account as this is the only allowed scope for secrets.
Finally, create the secret using:
databricks secrets put --scope <name of scope> --key DataLakeStore
In the notebook, the secret can then be used to connect to ADLS using the following configuration:
spark.conf.set("fs.azure.account.key.datalakeaccount.dfs.core.windows.net", dbutils.secrets.get(scope = "<name of scope>", key = "DataLakeStore"))
spark.conf.set("fs.azure.createRemoteFileSystemDuringInitialization", "true")
dbutils.fs.ls("abfss://datalakeaccount.dfs.core.windows.net/")
spark.conf.set("fs.azure.createRemoteFileSystemDuringInitialization", "false")
The file could then be loaded into a Spark dataframe via:
df = sqlContext
.read
.format('csv')
.options(header='true', inferSchema='true', delimiter='\\t')
.load('abfss://datalakeaccount@datalakeaccount.dfs.core.windows.net/Raw/data.tsv')
Unfortunately, though the pandas read function does work in Databricks, we found that it does not work correctly with external storage. In order to get a pandas dataframe you can use:
pandas_df = df.toPandas()
Alongside the setting:
spark.conf.set("spark.sql.execution.arrow.enabled", "true")
This setting enables the use of the Apache Arrow data format. This allows the data to be converted into Arrow format, which can then be converted via zero allocation pyarrow to convert into a Pandas dataframe. Without this setting, each row must be individually pickled (serialized) and converted to Pandas which is extremely inefficient. However, we still ran into problems when trying to convert back into a spark array to write back out to the file system around the types of the various columns. At this point, I was beginning to suspect that it was going to make sense to use PySpark instead of Pandas for the data processing.
Parallel processing using Spark
PySpark is the Python API used to access the Spark engine. Once data is loaded into a Spark dataframe, Spark processing can be used via this API for manipulation and transformations. It also helps you to access and manipulate the RDDs (Resilient Distributed Databases) which Spark is based on. An RDD is an immutable collection of elements of data, which is distributed across the cluster.
Alongside the inefficiency in converting to and from Pandas, there is another key motivator for the switch to PySpark. Spark has in-built optimisation which means that when you are working with large dataframes jobs are automatically partitioned and run in parallel (over the cores that you have available). This is implemented via the RDDs mentioned above, in order to distribute the processing. This parallelization allows you to take advantage of the autoscale features in Databricks. When you set up a (job or interactive) Databricks cluster you have the option to turn on autoscale, which will allow the cluster to scale according to workload. If you combine this with the parallel processing which is built into Spark you may see a large boost to performance. (A word of warning, the autoscale times are along the lines of the cluster spin up/down times so you won't see much of a boost for short running jobs.). So, with this in mind, I started on the conversion.
There were a few things which needed to be done to get the data into the correct format. The first was to replace certain values with null. The original processing was done using: na_values=['Na','NaN','None','_','NONE'] as an argument of the Pandas read function. There is no way to specify null values on import in PySpark so instead we did the following:
from pyspark.sql.functions import col, when
def blank_as_null(x):
return when(((col(x) != "None") & (col(x) != "NONE") & (col(x) != "_") & (col(x) != "Na") & (col(x) != "NaN")), col(x)).otherwise(None)
columns = df.columns
column_types = dict(df.dtypes)
for column in columns:
if column_types[column] == 'string':
df = df.withColumn(column, blank_as_null(column))
The other problem was that some of our column headings contained fullstops, which is not supported by PySpark. Therefore, we used the following function to escape the column headings:
def escape_columns(df):
for col in df.columns:
if '.' in col:
df = df.withColumnRenamed(col, col.replace('.','_'))
return df
df = escape_columns(df)
Once these changes had been made we were ready to start processing the data. For example, one of the steps in the ETL process was to one hot encode the string values data in order for it to be run through an ML model. This means to create a sparse numerical matrix which represents categorical data. This was originally done using the Pandas get_dummies function, which applied the following transformation:
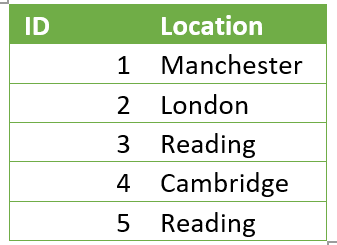
Turned into:
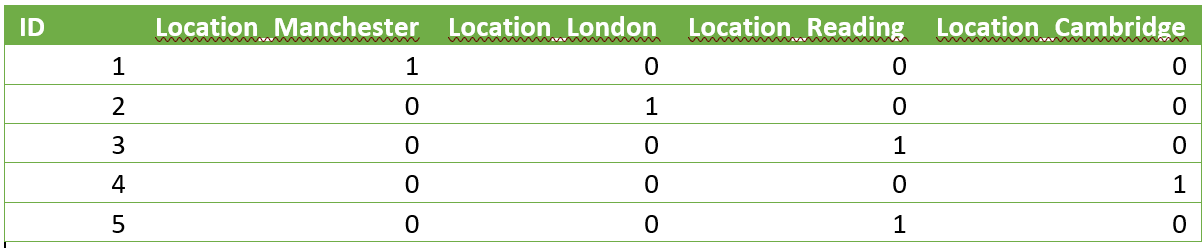
To do the equivalent using PySpark, we used the following:
def one_hot(df, feature):
feature_values_df = df.select(feature).distinct()
feature_values = [str(row[feature]) for row in feature_values_df.collect()]
for feature_val in feature_values:
df = df.withColumn(feature+"_"+feature_val, when(df[feature] == feature_val, 1).otherwise(0))
df= df.drop(feature)
return df
To perform the one hot encoding on the necessary columns over the dataframe.
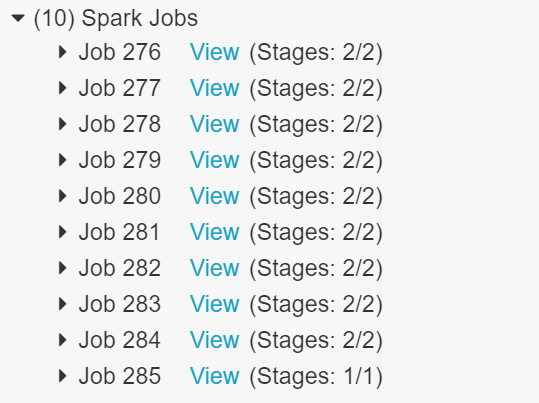
Once we had switched the ETL process over to use Spark we could see the splitting up of different dataframe operations into multiple jobs. This work can then be split up over the number of workers in your Databricks cluster.
Another parallel processing option which I think is worth mentioning is the multiprocessing Python library. This works with either Pandas or Spark and can be used to explicitly split tasks over multiple workers. You set up a thread pool using the number of available workers:
import multiprocessing as mp`
num_cores = mp.cpu_count()
pool = mp.Pool(num_cores)
Another warning: this will evaluate the number of workers at the time, and set this to the maximum number of nodes to use. Therefore, if you are using the autoscale functionality it will not recognise that the number of workers has increased. You can then use the pool.map(function_name, list) function to split up jobs across those cores. Here list is the list of individual values to run through function_name concurrently. This form of parallel processing can be especially useful when running the same time consuming tasks multiple times.
Once you have performed your ETL processing (using whatever parallelisation techniques you think are necessary), you can output the data back into ASDL Gen2 using:
df_num.write.format('csv')
.mode('overwrite')
.option('header', 'true')
.save('abfss://datalakeaccount@datalakeaccount.dfs.core.windows.net/Processed/Data')
This will output the data as many partitioned CSVs. These CSVs can then be used as a source in ADF (which can read in the data from the CSVs and combine into one dataset) and more processing can be carried out (via ADF Mapping Data Flow, another Notebook, etc!).
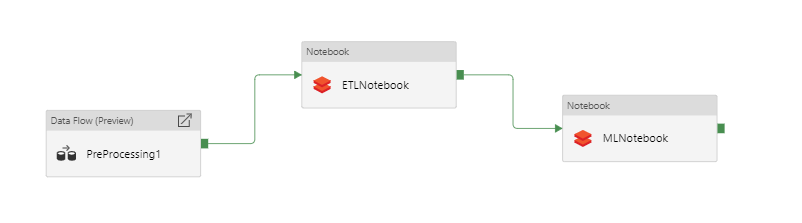
Overall, data manipulation via Databricks Notebooks can be combined with various other processing activities in ADF to build up complex and scalable ETL pipelines. We think this is a very exciting prospect and can't wait to hear what's next in this space!